For thousands of years, humans have been trying to understand the diversity of life on earth. Many of our systems of classification are based on bodies (phenotypes or morphologies). For example, as a paleoethnobotanist, I study tiny fragments of ancient plants that were left behind by people at archaeological sites – mostly in the garbage. I use the shape and size of ancient seeds to discover which species they came from, and then to describe variation in populations, especially of crop plants. The bodies of ancient organisms can tell us a lot about the origins and evolution of biodiversity. Whole fields are based on this premise: paleontologists study fossils; bioarchaeologists, paleoethnobotanists and zooarchaeologists study ancient remains of humans, plants, and animals recovered from archaeological sites. Up until recently, extinct forms were scientifically priceless, but held no economic value for agriculture or medicine. De-extinction was a topic for science fiction.

Though usually remembered for its depiction of dino de-extinction, Dr. Ellie Sattler’s reaction to this leaf (“Alan, this species has been extinct since the Cretaceous”) at the beginning of Jurassic Park’s most iconic scene implies that the thinking-machine-super-computer had also managed to ressurect lost plant species. Sadly, this never comes up again in any of the many Jurassic Park films.
But we’re currently witnessing a revolution in the study of ancient biodiversity: genomics has cannonballed into archaeology and made an epic splash. With this methodological sea change, we have an opportunity to address fundamental questions that were unanswerable with older methods, but we also face new ethical questions. Much ink has already been spilled announcing the arrival of the genomics revolution in the study of ancient human DNA. There has been an explosion of research in the past decade, enabled by next-generation sequencing and breakthroughs in methods for separating out damaged fragments of ancient DNA from modern contaminants. Since we humans are naturally quite interested in ourselves, it is not surprising that these new techniques have been applied earlier and more widely to the study of human evolution than to that of plants and animals. Already, ancient genomic approaches have been used to better understand the sex lives of ancient humans, Neanderthals, and Denisovans (spoiler alert: they were all having sex with each other, and it was adaptive!), to test old hypotheses about the demographic effects of Neolithic Revolution, and to understand the evolution of specific human adaptations, such as the ability to digest lactase in adulthood and low hemoglobin levels in high altitude populations.
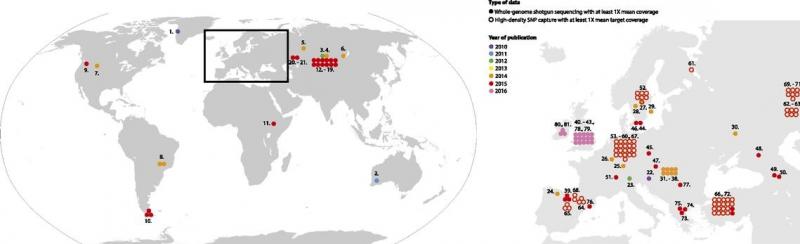
From Slatkin and Racimo, 2016, this figure shows the rapid increase in recovery of ancient human genomes between 2010 and 2016. In the past two years, this trend has continued and intensified.
These studies are revolutionizing how we understand human evolution and history, but they have also spawned very serious problems. Renowned human geneticist David Reich recently caused a snafu with an editorial in the New York Times conflating populations (in the biological sense of the word) with races and arguing that it is “no longer possible to ignore average genetic differences among “races.”” (Click here for more on this controversy). Meanwhile, white supremacists are chugging milk to demonstrate their European ancestry, in an homage to one of the studies cited above. Researchers have also raised concerns about the (sometimes) wanton destruction of ancient human remains for aDNA extraction, and how irreplaceable and rare specimens are being hoarded by a few elite institutions. These concerns are about to become major problems in the study of ancient plant DNA, too, because…
Screenshot of white supremicists chugging milk at a protest in New York City, as reported by Amy Harmon.
A revolution is also underway in biotechnology. Maybe you’ve heard about CRISPR/Cas9 and thought “That’s just too much acronym for me to process today,” but please bear with me. This discovery is already transforming our world – and especially our food. Previous methods of genetic modification involved infiltrating or blasting cell nuclei with packages of modified genes and hoping they incorporated properly (1,2). Because the new genes incorporated more or less randomly into the target organism’s genome, biotechnologists had to rely on a brute force evaluation of thousands of modified plants or animals to identify a few in which the new genes had been properly incorporated, yielding the desired new trait. Making multiple targeted changes using these methods was extremely labor and time intensive. CRISPR/Cas9, on the other hand, is a gene editing tool that allows modified or novel genes to be inserted into the target genome at precisely the right place.
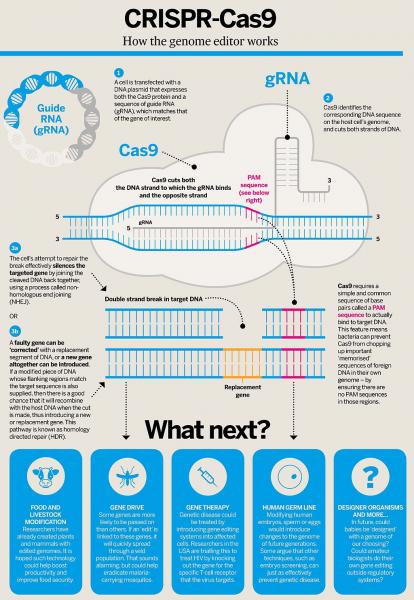
Schematic of CRISPR/Cas9 system, courtesy of Wikimedia Commons.
This method makes use of two defense mechanisms that bacteria use to fight off viruses. CRISPR (clustered regularly interspaced short palindromic repeats, in case you were wondering) is a piece of RNA created by bacteria to help them recognize viruses that they have previously encountered. After recognizing a virus, bacteria deploy Cas9, an enzyme that destroys the virus by cutting apart its genome (you’ll sometimes hear Cas9 referred to as “molecular scissors.” (Read more here!) Biotechnologists can now use modified CRISPR RNA to target specific places in the genome of the organism they are modifying, and Cas9 to insert the new gene at that place. This method is a game changer because it makes genetic modification much more precise and predictable. In the US, we have already seen our agricultural system transformed by genetically modified crops created using much clunkier methods. Not only is CRISPR/Cas9 gene editing an extremely efficient method, the USDA has officially declined to regulate CRISPR/Cas9-edited crops by reframing this technology as a “plant breeding innovation.” There are already five CRISPR-edited products being developed for the market, including soybeans, camelina, lawn grass, maize, and mushrooms.
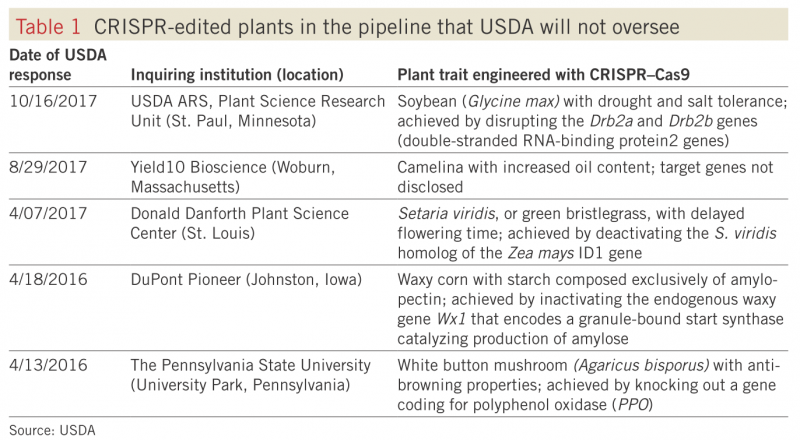

From Walz 2018, reused with permission from Springer.
What do these two scientific breakthroughs – ancient genomics and CRISPR/Cas9 gene editing – have to do with ancient garbage? That’s the million-dollar question.
This year, two reviews appeared, one in Nature: Plants, and the other in Frontiers in Plant Science, both arguing that ancient plant remains have just become a source of valuable information for the biotech and crop breeding industries. Their argument is based on two points:
- We are suddenly able to recover millennia worth of lost agrobiodiversity from around the world.
- We have simultaneously developed the tools to directly incorporate useful variation from ancient plants into modern crops.
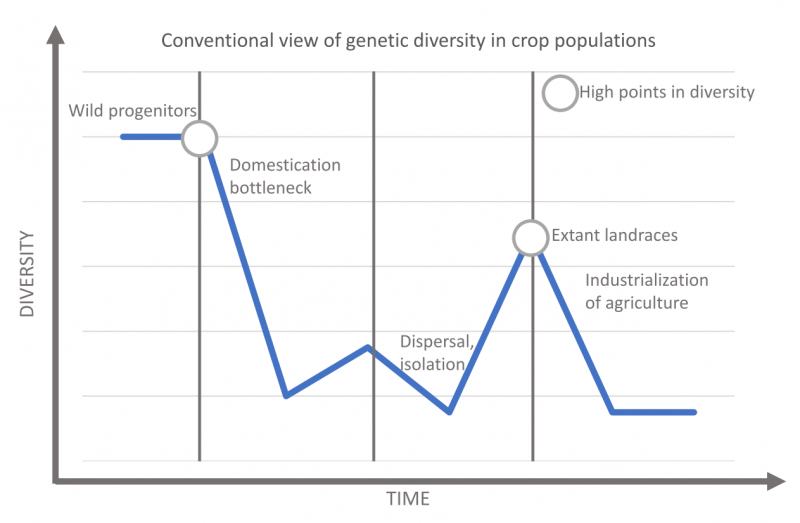
The conventional view of the evolution of agrobiodiveristy. After Allaby et al. 2018, Fig 1.
To start with the first point, why would ag industries care about the variation contained in ancient crop genomes? Well, first take a gander at my schematic of the “conventional view” of the relationship between genetic diversity and domestication. In this view, there is a steep drop in diversity, called the domestication bottleneck, when a small subset of a large wild population is brought under cultivation. For most crops, this happened thousands of years ago. From there, development of different varieties led to increasing biodiversity within the crop population. We call the results of this diversification landraces, which are regionally specific crop varieties, often adapted to local tastes or environmental conditions. When crops were taken into new regions this could have led to more bottlenecks, especially if dispersal was followed by isolation. Then diversification would have begun again in multiple regions, and sometimes these distinct populations came back together through trade and migration, leading to a huge amount of agrobiodiversity worldwide.
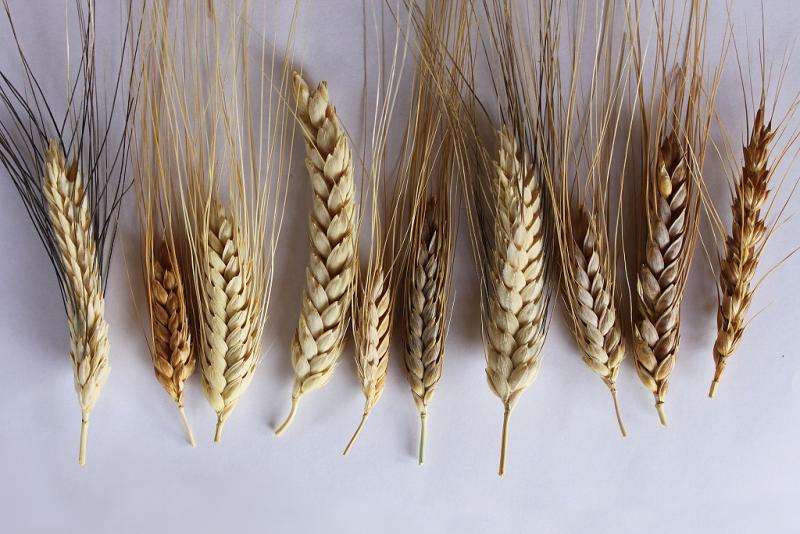

Wheat landraces collected in Turkey. Image by Alexei Morgounov/CIMMYT.
Finally, we have a known bottleneck in agrobiodiversity that started with plantation economies and colonialism and culminated in the industrialization of agriculture during the 20th century. During this process, seed selection and crop breeding moved off of farms and most landraces fell out of cultivation, although many are still preserved in seed banks and herbaria. Today, the overwhelming majority of seed planted globally comes from crop breeders or biotech companies. They only produce a relatively limited number of “elite” varieties that have usually been bred for yield. Pant breeders and biotechnologists have always known that landraces and the wild progenitors of crops are diversity high points, and sources of useful genetic variation for crop improvement. They can use the relative diversity of these populations to breed or engineer useful traits into commercial varieties – things like drought and virus resistance, particular tastes, or different nutritional properties.
But recent insights from ancient crop genomes have called the assumptions of the conventional model of crop evolution into question. Ancient genomic data for three crops (maize, barley, and sorghum) seems to indicate that there was no domestication bottleneck. Instead of a sharp drop in diversity immediately after domestication, these crops experienced a gradual loss in genetic diversity over the course of thousands of years. One important implication of this research is that the genomes of ancient crops may hold useful traits that were lost long before colonizing botanists started to fill seed banks with landraces. In other words, the amazing variety of crops that were grown around the world when the Age of Exploration began may not actually be a high point in diversity. The high point may instead have come thousands of years ago, and there is likely a unique evolutionary history for every crop.
This would all be academic if it weren’t for simultaneous breakthroughs in gene editing. You can’t (usually) get ancient seeds to germinate, so however much valuable diversity these lost varieties hold, they would have been useless to modern plant breeders before genetic modification. But now, using gene editing, it is possible to reverse engineer living crops to replicate desirable aspects of ancient varieties.
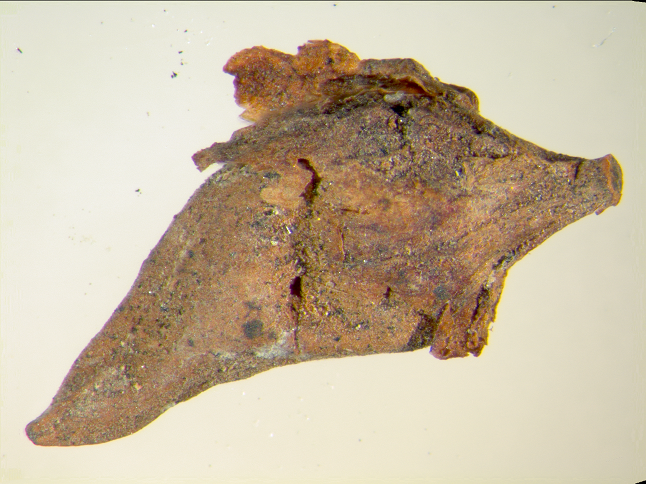

My favorite lost crop: ~2,500 year old dessicated erect knotweed fruit from Cold Oak rockshelter, KY. Image by Natalie G. Mueller
How you feel right now is probably largely determined by your opinions about genetically modified crops in general. Personally, I find the idea of de-extincting lost crops using gene editing, frankly AWESOME. But I know enough about the history of GM crops to guess that research along these lines will not be driven by my desire to have a historically accurate garden. Instead, ancient genomes will be mined for profitable traits, and what was once the cultural heritage of particular communities will become extremely valuable private property through the alchemy of formal breeding and genetic modification. The archaeobotanical collections that could make this kind of research possible are extremely rare, fragile, and difficult to recover. They are a limited and non-renewable resource. Curators, archaeologists, and descendent communities should get started now developing ethical and legal frameworks to guide the use of archaeobotanical specimens, before the application of ancient genomic studies expands to plant breeding and biotechnology.
Sources
Allaby, R. G., Ware, R. L., & Kistler, L. A re-evaluation of the domestication bottleneck from archaeogenomic evidence. Evolutionary Applications, 0(0). doi:doi:10.1111/eva.12680
Di Donato, A., Filippone, E., Ercolano, M. R., & Frusciante, L. (2018). Genome Sequencing of Ancient Plant Remains: Findings, Uses and Potential Applications for the Study and Improvement of Modern Crops. Frontiers in plant science, 9(441). doi:10.3389/fpls.2018.00441
Estrada, O., Breen, J., Richards, S. M., & Cooper, A. (2018). Ancient plant DNA in the genomic era. Nature plants, 4(7), 394-396. doi:10.1038/s41477-018-0187-9
Ficiciyan, A., Loos, J., Sievers-Glotzbach, S., & Tscharntke, T. (2018). More than Yield: Ecosystem Services of Traditional versus Modern Crop Varieties Revisited (Vol. 10).
Haak, W., Lazaridis, I., Patterson, N., Rohland, N., Mallick, S., Llamas, B., . . . Reich, D. (2015). Massive migration from the steppe was a source for Indo-European languages in Europe. Nature, 522, 207. doi:10.1038/nature14317
https://www.nature.com/articles/nature14317#supplementary-information
Huerta-Sánchez, E., Jin, X., Asan, Bianba, Z., Peter, B. M., Vinckenbosch, N., . . . Nielsen, R. (2014). Altitude adaptation in Tibetans caused by introgression of Denisovan-like DNA. Nature, 512, 194. doi:10.1038/nature13408
https://www.nature.com/articles/nature13408#supplementary-information
Itan, Y., Powell, A., Beaumont, M. A., Burger, J., & Thomas, M. G. (2009). The origins of lactase persistence in Europe. PLoS computational biology, 5(8), e1000491.
Jaganathan, D., Ramasamy, K., Sellamuthu, G., Jayabalan, S., & Venkataraman, G. (2018). CRISPR for Crop Improvement: An Update Review. Frontiers in plant science, 9, 985-985. doi:10.3389/fpls.2018.00985
Kuhlwilm, M., Gronau, I., Hubisz, M. J., de Filippo, C., Prado-Martinez, J., Kircher, M., . . . Castellano, S. (2016). Ancient gene flow from early modern humans into Eastern Neanderthals. Nature, 530, 429. doi:10.1038/nature16544
https://www.nature.com/articles/nature16544#supplementary-information
Llamas, B., Willerslev, E., & Orlando, L. (2017). Human evolution: a tale from ancient genomes. Philosophical Transactions of the Royal Society B: Biological Sciences, 372(1713). doi:10.1098/rstb.2015.0484
Makarewicz, C., Marom, N., & Bar-Oz, G. (2017). Ensure equal access to ancient DNA. Nature, 548, 158. doi:10.1038/548158a
Prendergast, M. E., & Sawchuk, E. (2018). Boots on the ground in Africa’s ancient DNA ‘revolution’: archaeological perspectives on ethics and best practices. Antiquity, 92(363), 803-815. doi:10.15184/aqy.2018.70
Racimo, F., Sankararaman, S., Nielsen, R., & Huerta-Sánchez, E. (2015). Evidence for archaic adaptive introgression in humans. Nature Reviews Genetics, 16, 359. doi:10.1038/nrg3936
https://www.nature.com/articles/nrg3936#supplementary-information
Seguin-Orlando, A., Korneliussen, T. S., Sikora, M., Malaspinas, A.-S., Manica, A., Moltke, I., . . . Willerslev, E. (2014). Genomic structure in Europeans dating back at least 36,200 years. Science, 346(6213), 1113-1118. doi:10.1126/science.aaa0114
Slatkin, M., & Racimo, F. (2016). Ancient DNA and human history. Proceedings of the National Academy of Sciences, 113(23), 6380-6387. doi:10.1073/pnas.1524306113
Waltz, E. (2018). With a free pass, CRISPR-edited plants reach market in record time. Nature Biotechnology, 36, 6. doi:10.1038/nbt0118-6b
